Announcements
- May 4: The final exam is on-line and can be downloaded from this link.
- May 3: The code to compute global alignment, GlobalAlign(v, w, scorematrix, indel), had a bug. An updated, corrected version has been posted in the Lecture 16 notebook. The lecture notes have also been upadted. You should also notice that the code for LocalAlign(v, w, scorematrix, indel) appears in a non-printed cell of the Lecture 17 notebook.
- April 30: The final exam study session will be Tuesday, May 1 from 5-7pm in FB007.
- April 26: Problem Set #5 is now graded. Please speak to or email Maya if you have any issues related to your course grades.
- April 24: Problem Set #4 is graded.
- April 17: We will do the last problem set #5 together in class on April 19. It will be due at the end of class, the late policy will take effect immediately after.
- April 11: I will be unable to make my office hours this morning. If you have questions, please email me. -- Maya
- March 29: There are a few typos in Problem Set #4. In Problem #2, the first example string should match the second, "TAGACTAAG". Problem #3 should say: "In the space below give a recurrence relation...". An updated version of the problem set has been posted.
- March 26: Problem Set #4 is online and due on 4/5.
- March 22: Grades for the midterm are online
- March 20: Grades for Problem Set #2 are online.
- March 8: The Midterm examination can be downloaded starting at 9:30am. Your final version must be uploaded before 10:50am.
- March 5: There was an error in the k-mer text file referenced in Question 1 & 2 of PS3. If you have already downloaded it, please re-download. Also, clarifications to some of the questions in PS3 have been made. A revised version of Problem Set #3 has been posted.
- March 5: If you need to reschedule your midterm exam, you must do so by the end of class tomorrow (Tuesday, March 6). Please email Maya (mnajarian@cs.unc.edu) with your excuse and a time period during the week of March 10-17 when you can take the midterm.
- March 3: I added several missing links to the schedule for Lecture 14 (slides and notebook). Also Problem Set #3 is now online and is due on 3/22. I would suggest that you look at it before the midterm.
- February 23: There was a mistake the example in Question 5 of PS2. There are actually 6 index combinations for the subsequence, instead of 2. A revised version of Problem Set #2 has been posted.
- February 20: I will be unable to make the first hour of my office hours today from 2pm-3pm. I will hold additional makeup hours next week. -- Leonard
- February 15: Problem Set #2 is online and due on 3/1.
- February 13: In problem 4 of PS #1, "chromosome 1" should be changed to "complete genome". You should used the sequence with "complete genome" in its header. A revised version of Problem Set #1 has been posted.
- January 31: The Jupyter/Python tutorial will be held in SN011 on 2/1 from 5pm-6:30. Here is the notebook used in the tutorial.
- January 30: Problem Set #1 is online and due on 2/15. We still do not have a time and place for the Python/Jupyter study session. Stay tuned.
- January 25: The first problem set will be issued on Tuesday, January 30.
- January 18: Several students have asked me to provide versions of the lecture slides that can be easily annotated. So I have replaced the downloadable "slides" with pdf versions. If you prefer the html format used previously, you can download the "notebook" and within Jupyter notebook save the file in html format.
- January 17: Tomorrow's class has been cancelled. I will try to adjust things in the schedule to make up for it. Stay safe and enjoy the snow. Check here reguarly for announcements of the time and place of the Python study session.
- January 12: DON'T BUY THE TEXTBOOK! I did not realize that, since last spring's offering of this course, the textbook's price had changed so dramatically as shown in the following figure from camelcamelcamel.com:
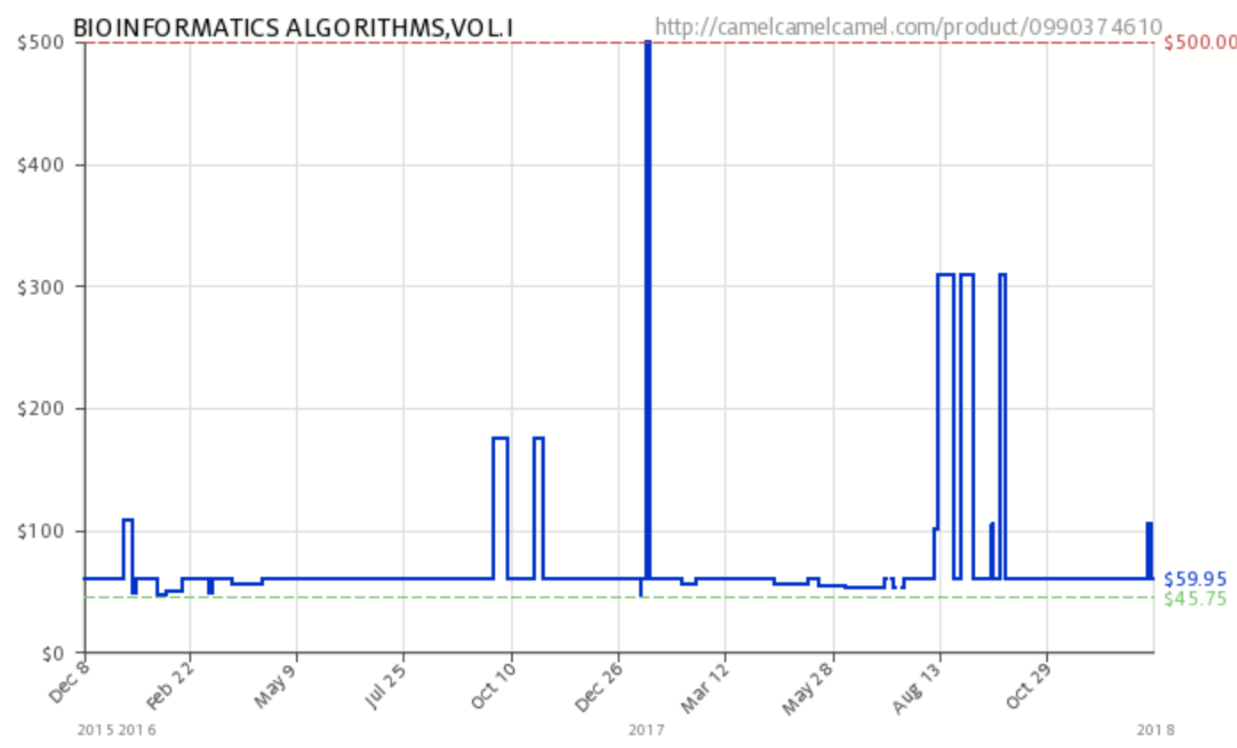 Just yesterday the price was over $100. Considerably more than the suggested $45.75 or average of $59.95. To me this looks like an Uber "surge pricing" model designed to gouge students. I don't want to be any party to this, so don't buy the book for now. I have notes from a different book that I might switch to, but let's all work from my on-line notes to start and look to buy low and sell high. -Leonard
Just yesterday the price was over $100. Considerably more than the suggested $45.75 or average of $59.95. To me this looks like an Uber "surge pricing" model designed to gouge students. I don't want to be any party to this, so don't buy the book for now. I have notes from a different book that I might switch to, but let's all work from my on-line notes to start and look to buy low and sell high. -Leonard - January 11: First class meeting in SN014. See you there
Course Description
Computational methods are fueling a revolution in the biological sciences. Computers are already nearly as indispensable as microscopes for analyzing and interpreting biological data. As a result, two new multidisciplinary fields, bioinformatics and computational biology, have emerged. This course will explore the computational methods and algorithmic principles driving this revolution. It will cover basic topics in molecular biology, genetics, and proteomics. The course also addresses basic computational theory and algorithms including asymptotic notation, recursion, divide-and-conquer approaches, graph algorithms, dynamic programming, and greedy algorithms. These fundamental concepts from computer science will be taught within the context of motivating problems drawn from contemporary biology. Example biological topics include sequence alignment, motif finding, gene rearrangement, DNA sequencing, protein peptide sequencing, phylogeny, and gene expression analysis.
This course is suitable for both computer science and biology students at both undergraduate and graduate levels. Students who wish to take this course should have some programming experience in a modern programming language. Knowledge of data structures, algorithm design, and biology is helpful but not required. There will be 5 problem sets each with short programming assignments (each worth 10%), a midterm (worth 20%), and a final exam (worth 30%).
A syllabus for this offering of Comp555 can be downloaded from here.
Book, Course Information, and Prerequisites
Here is the book, which I will be supplementing with additional materials:
 Bioinformatics Algorithms: An Active Learning Approach, Vol 1
Bioinformatics Algorithms: An Active Learning Approach, Vol 1
by Phillip Compeau and Pavel Pevzner
Active Learning Publishers © 2014, ISBN: 0990374610.
See the annoucement from January 12. Hold off buying a textbook for now. There is another book that I can use, and I might try doing this years course using entirely on-line materials.
| Credit Hours: | 3 |
| Location: | SN014 |
| Time: | TTh 9:30-10:45 |
| URL: | http://www.csbio.unc.edu/mcmillan/?run=Courses.Comp555S18 |
| Prerequisites: | COMP 410, Math 381, or equivalents |
Course Instructors
| Instructor: | Leonard McMillan |  |
| Office: | SN316 | |
| email: | mcmillan@cs.unc.edu | |
| Office Hours: | Tuesdays 2pm-4pm |
| TA: | Maya Najarian |  |
| Office: | SN312 | |
| email: | mnajarian@cs.unc.edu | |
| Office Hours: | T 4-5, W 10-11 |
Schedule
| Date | Topic | Homework |
| January 11 | Introduction (slides, notebook) | |
| January 16 | Exploring a Genome (slides, notebook) | (Video Parts 1 & 2) Reading: Chapter 1 pp 3-14 |
| January 28 | Class cancelled due to inclement weather | |
| January 23 | Exploring a Genome (Continued) (slides, notebook) | (Video Parts 3, 4, & 5) Reading: Chapter 1 pp 14-45 |
| January 25 | Finding Patterns in DNA (slides, notebook) | (Video Parts 1 & 2) Reading: Chapter 3 pp 83-92 |
| January 30 | Searching for Motifs (slides, notebook) | (Video Parts 3, 4, 5 & 6) Reading: Chapter 3 pp 93-127 Problem Set #1 Assigned |
| February 1 | Protein Sequences and Antibotics (slides, notebook) |
(Video Parts 1,2, & 3) |
| February 1 | Crash course in Jupyter, Python, and Rosalind (notebook) | Link: Rosalind |
| February 6 | Inferring Protein Sequences from Fragments (slides, notebook) |
(Video Parts 4, 5, & 6) |
| February 8 | Scaling up Peptide Sequencing (slides, notebook) | (Video Parts 7, 8, 9, & 10) Reading: Chapter 2 pp 66-80 |
| February 13 | Assembling a Genome (slides, notebook) | (Video Parts 1, 2, 3 & 4) Reading: Chapter 4 pp 129-152 |
| February 15 | Path Finding in Graphs (slides, notebook) | (Video Parts 4, 5, 6, 7, & 8) Reading: Chapter 4 pp 153-187 Problem Set #2 Assigned |
| February 20 | The Realities of Genome Assembly (slides, notebook) | (Video Parts 9, 10, 11, & 12) |
| February 22 | Combinatorial Pattern Matching (slides, notebook) | (Video Parts 1-3) |
| February 27 | Suffix Trees and BWTs (slides, notebook) | (Video Parts 4-9) |
| March 1 | Multi-String BWTs (slides, notebook) | (Not in Book) Problem Set #3 Assigned |
| March 6 | Comparing Sequences (slides, notebook) | (Video Parts 1, 2, 3, & 4) Reading: Chapter 5 pp 189-199 |
| March 8 | Midterm, Open book, open notes, online (covers to Lecture 13, 2/27) | |
| March 13 | No Class (Spring Break) | |
| March 15 | ||
| March 20 | Comparing Sequences (cont) (slides, notebook) | |
| March 22 | Go over Midterm Sequence Alignment (slides, notebook) |
(Video Parts 6, 7, & 8) Problem Set #3 due Problem Set #4 Assigned |
| March 27 | Advanced Sequence Alignment (slides, notebook) | (Video Parts 10, & 11) Reading: Chapter 5 pp 230-258 |
| March 29 | Generalized Dynamic Programming (slides, notebook) | (Video Part 5) Reading: Chapter 5 pp 201-206 |
| April 3 | Divide and Conquer Algorithms (slides, notebook) | (Video Part 9) Reading: Chapter 5 pp 236-243 |
| April 5 | Hidden Markov Models (slides, notebook) | (Video Parts 1, 2, 3, 4, & 5) |
| April 10 | Hidden Markov Models - Continued (slides, notebook) | (Video Parts 6-10) |
| April 12 | Genome Rearrangements (slides, notebook) | (Video Parts 1-3) |
| April 17 | Genome Rearrangements (cont) (slides, notebook) | (Video Parts 4-5) |
| April 18 | Genome Rearrangements (cont) (slides, notebook) | Problem Set #5 Assigned Due by end of Class |
| April 24 | Randomized Algorithms (slides, notebook) | (Not in book) |
| April 26 | Randomized Algorithms (continued) (slides, notebook) | (Not in book) |
| May 1 | Final Study Session in FB007 5:00pm-7:00pm | |
| Friday, May 4 | Final Exam (SN014) 8:00am-11:00am | |
Resources
- Download the package management system Anaconda. It will greatly simplify working with both Python 2.7 and Jupyter.
- After installing Anaconda, follow the instructions for installing Jupyter Notebook.