UNC Systems Genetics
| Home |
| Genetics | G3 Multiparental Populations (2017) |
| Genetics | G3 Special Issue (2012) |
| CC Mice |
| Information |
| Available Strains |
| Conditions of Use |
| CC Viewer |
| Locus Probabilities |
| Founder Probabilities |
| CC Genomes |
| CC Gene Expression |
| Genotypes |
| Platform Overviews |
| Imputed |
| MDA Tools |
| Mouse Phylogeny Viewer |
| Data |
| Genetics 2017 |
| Genetics 2012 |
| Publications |
| Order Mice |
| Contact Us |
The SGCF no longer offers MUGA genotyping service. Please contact GeneSeek at bpejsar@neogen.com
MegaMUGA genotyping
MegaMUGA is the second-generation Mouse Universal Genotyping Array. It extends MUGA from 7.5K to 77.8K markers, and includes all MUGA markers as a subset. There are three types of probes in MegaMUGA. In addition to traditional SNP probes, we have also introduced a second probe type for tracking known structural variants (insertions, deletions and duplications). A third probe type was designed to detect the presence of sequences present only in genetically engineered mice (Cre, Luciferase, etc).
The vast majority of MegaMUGA probes ascertain traditional biallelic SNPs. SNPs were selected to be distributed across the entire genome including the mitochondria and the Y chromosome with an average spacing of 33 Kb. For the autosomes, these probes were distributed as evenly as possible based on new linkage map for the mouse with a slight excess of probes in the telomeric regions to facilitate detection of recombination events in these regions.
Note that the MegaMUGA array is no longer available as of May 15, 2015. Please consider using the GigaMUGA array.
GigaMUGA genotyping
GigaMUGA is the third-generation Mouse Universal Genotyping Array. It extends MegaMUGA from 77.8K to approximately 150K markers, and includes all MUGA and most MegaMUGA markers as a subset.
MUGA genotyping
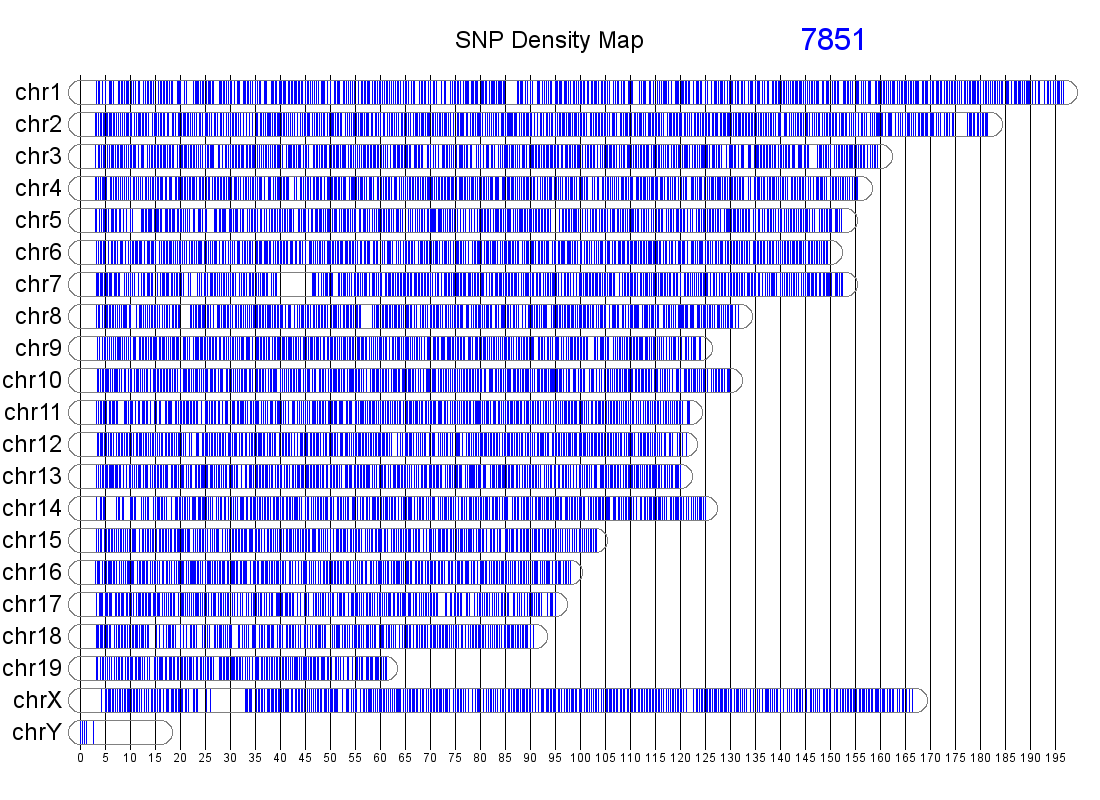
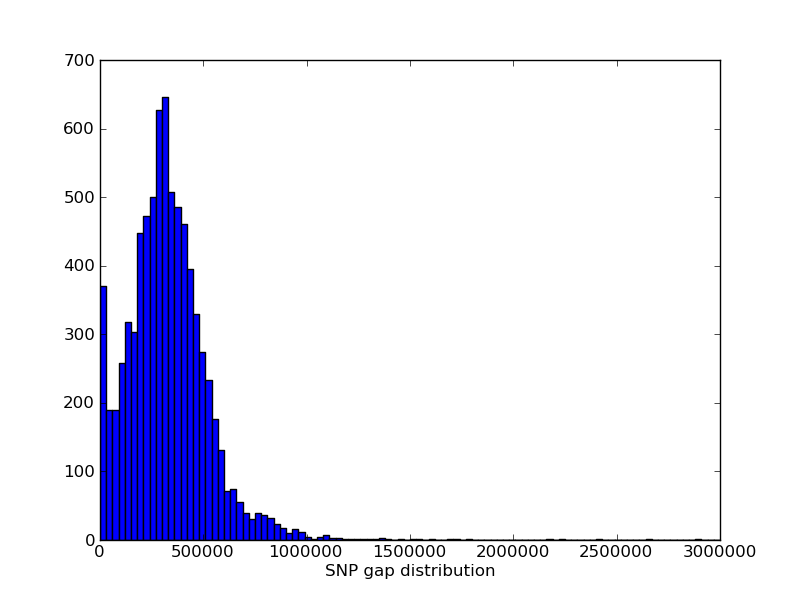
The genomic distribution of MUGA markers. The genomic position of each MUGA marker is shown on the left. The right figure illustrates the distribution of SNP spacing on MUGA. Click on either for a larger image.
MUGA is a custom designed 7851 SNP marker genotyping array built on the Illumina Infinium platform. SNP markers are distributed throughout the mouse genome with an average spacing of 325 kb (SD 191 kb). The markers were chosen to be maximally informative and maximally independent for the eight founder strains of the CC. MUGA has been used to genotype common laboratory strains, crosses, as well as wild mice.
Example MUGA genotypes can be downloaded here.
The MUGA array has been reissued and is useful for some applications.
MDA genotyping
MDA (no longer available through UNC) is an Affymetrix based genotyping platform with the following characteristics:
- Probes for more than 623,000 Single Nucleotide Polymorphisms (SNPs)
- 916,000 non-polymorphic probes distributed in functional, common CNV, and highly-conserved regions
The Mouse Diversity array captures the full range of genetic diversity present in classical laboratory mice, including classical and wild derived inbred strains. Other genotyping platforms have shortcomings when compared to MDA; they lack SNPs that discriminate between and within subspecies.
More information
For more information contact Darla Miller at: millerdr@med.unc.edu.